A mammalian methylation array for profiling methylation levels at conserved sequences | Nature Communications
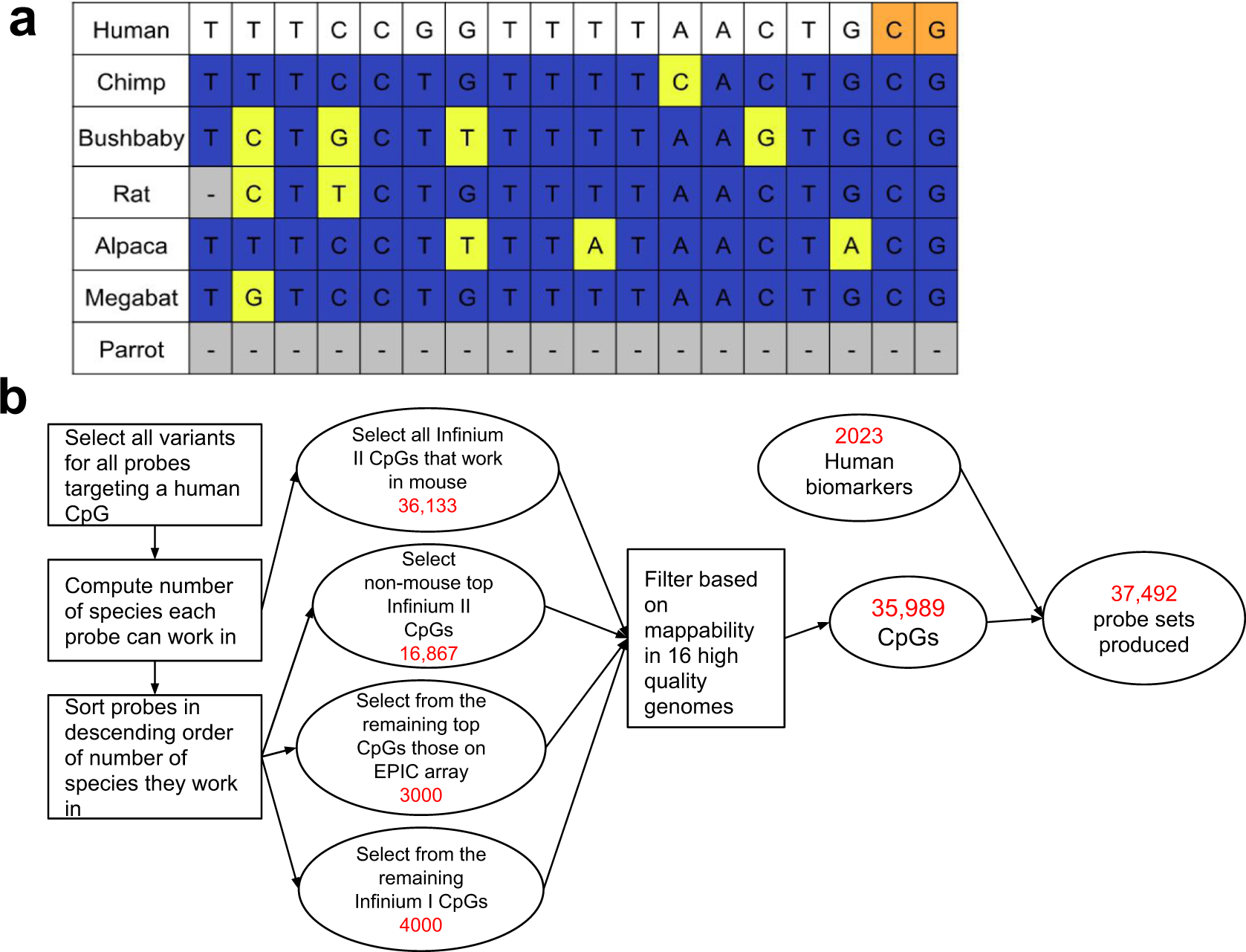
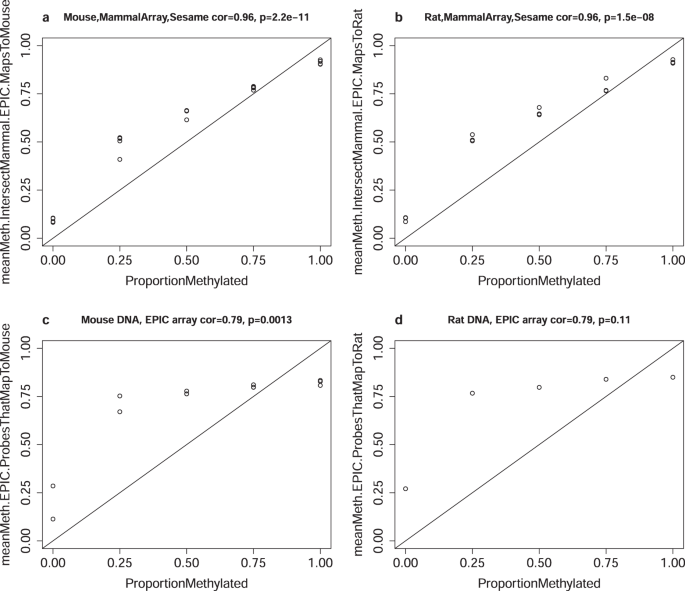
A mammalian methylation array for profiling methylation levels at conserved sequences | Nature Communications

Distribution of signal intensities and methylation β-values in mice and... | Download Scientific Diagram
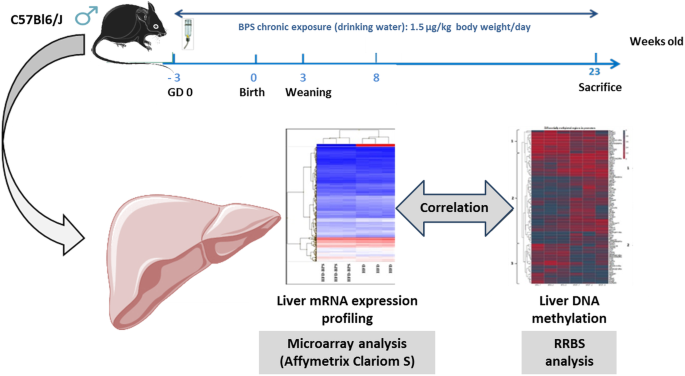
Hepatic transcriptome and DNA methylation patterns following perinatal and chronic BPS exposure in male mice | BMC Genomics | Full Text

Beadchip technology to detect DNA methylation in mouse faithfully recapitulates whole-genome bisulfite sequencing | Epigenomics
Comparative analysis of Illumina Mouse Methylation BeadChip and reduced-representation bisulfite sequencing for routine DNA meth

Comparative analysis of Illumina Mouse Methylation BeadChip and reduced-representation bisulfite sequencing for routine DNA methylation analysis - ScienceDirect

Comparative analysis of Illumina Mouse Methylation BeadChip and reduced-representation bisulfite sequencing for routine DNA methylation analysis - ScienceDirect
Comparative analysis of Illumina Mouse Methylation BeadChip and reduced-representation bisulfite sequencing for routine DNA meth

Mick Milsom on X: "Methylation analysis in mouse without the cost of whole genome bisulfite sequencing: check out Maxi's new analysis pipeline for methylation bead chip data. Congrats Maxi et al." /

Dynamic DNA methylation reveals novel cis-regulatory elements in mouse hematopoiesis - ScienceDirect

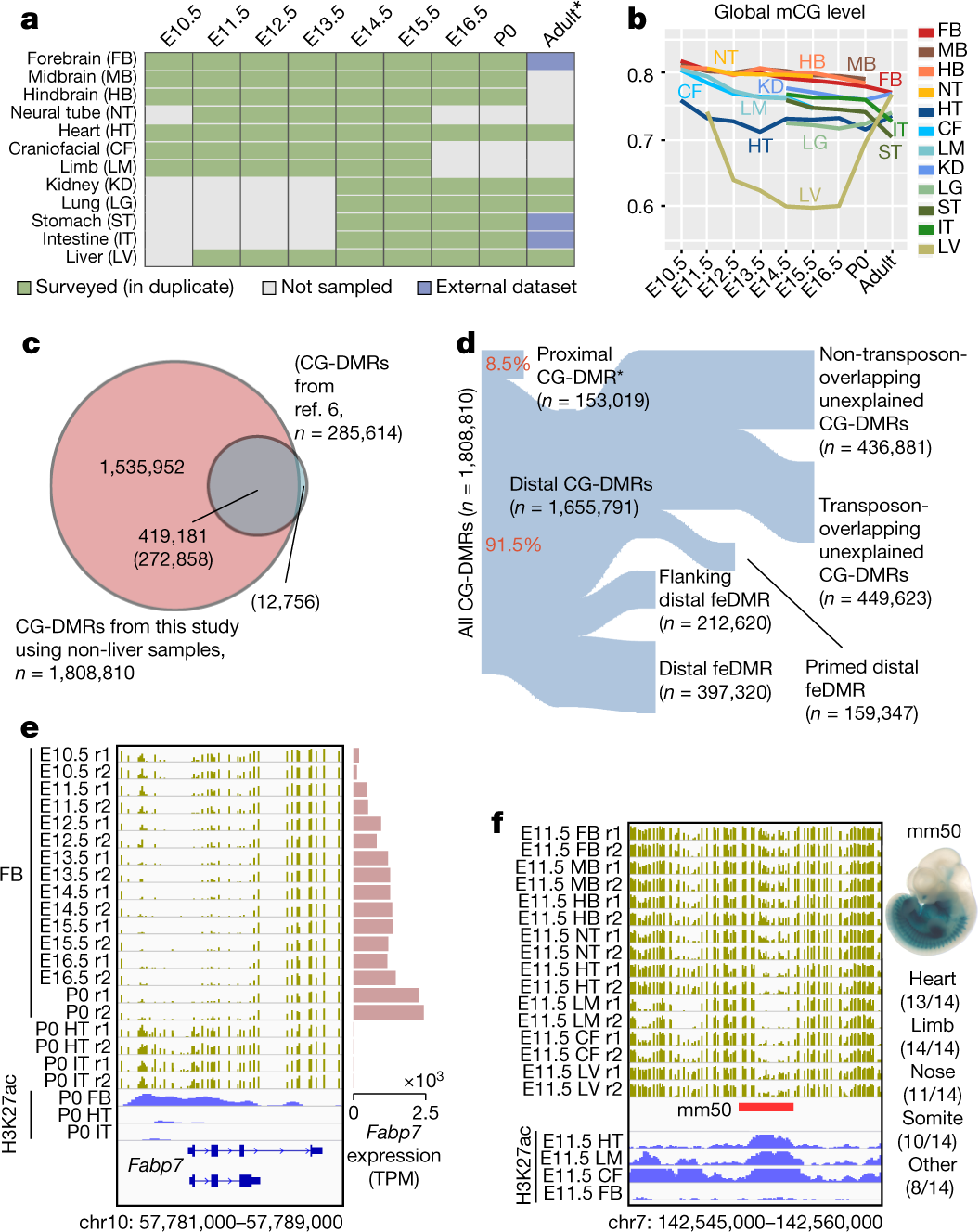
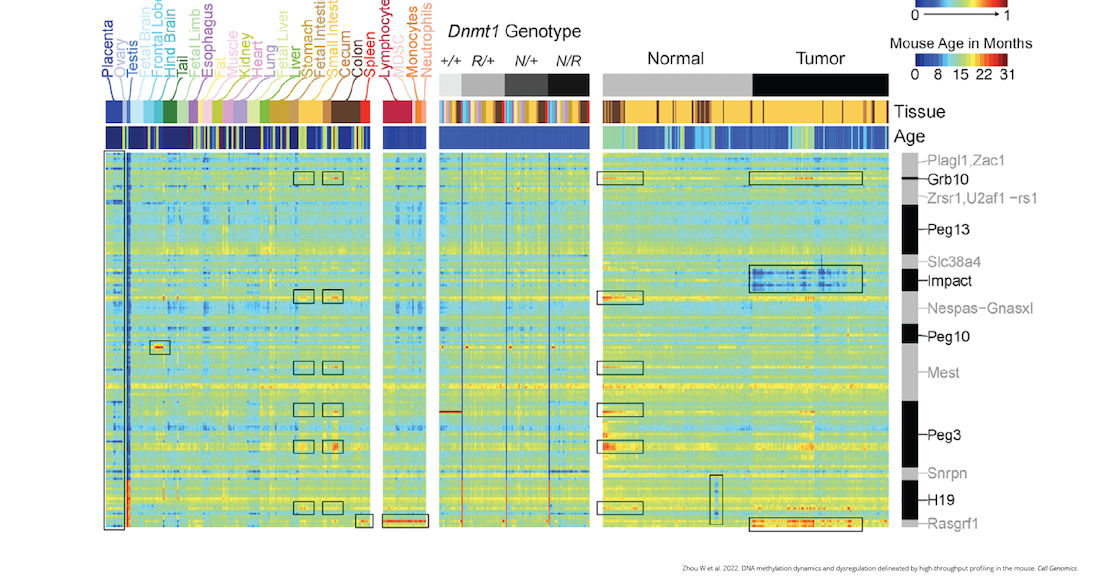
DNA methylation dynamics and dysregulation delineated by high-throughput profiling in the mouse - ScienceDirect
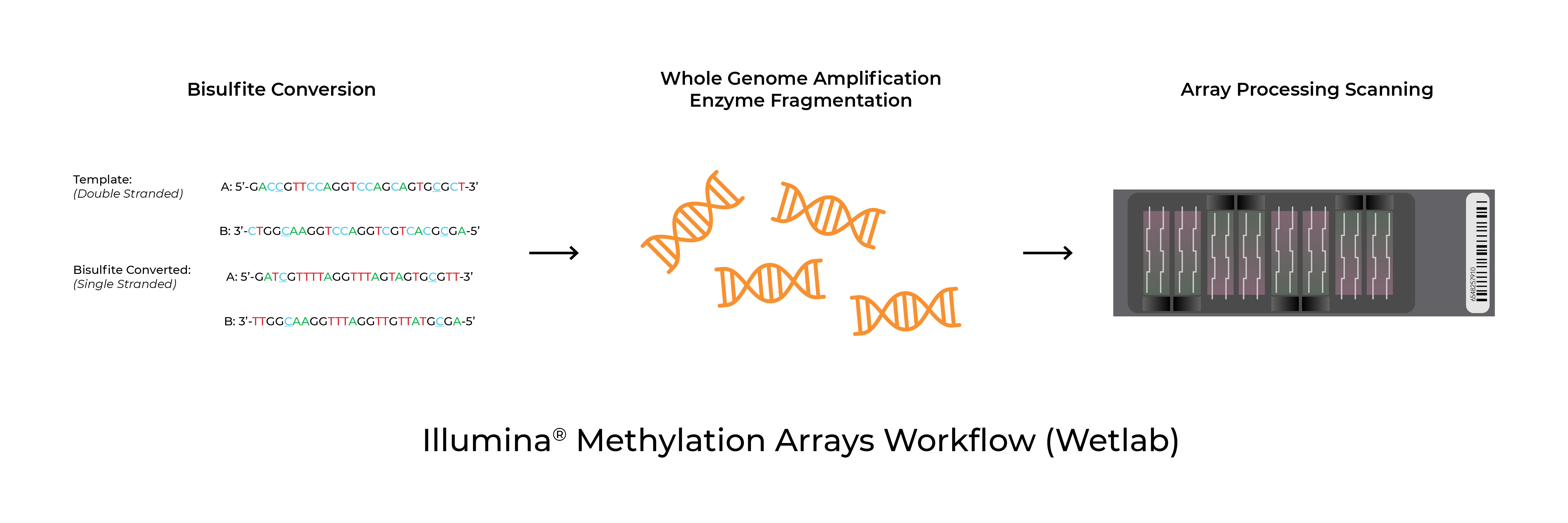
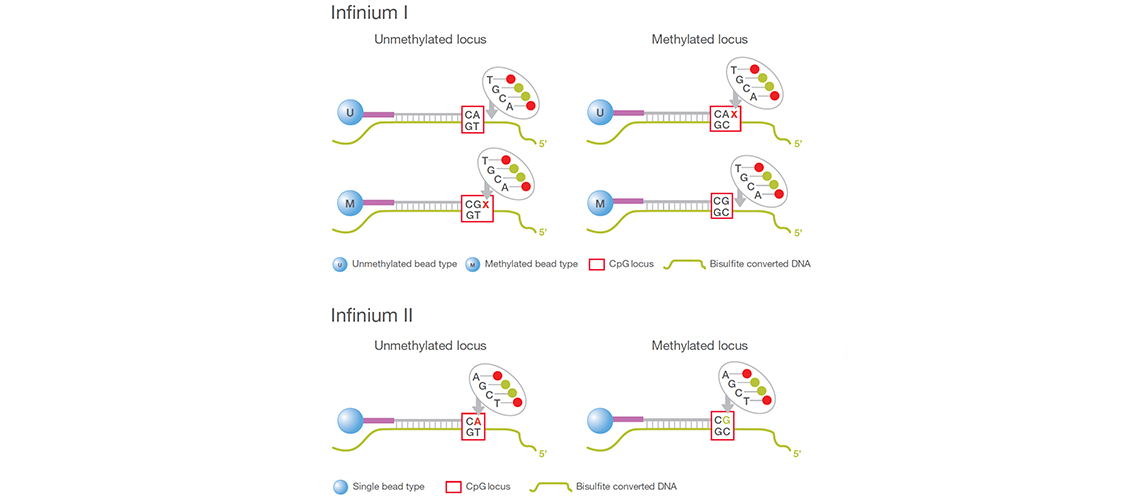
Bisulfite Conversion for Illumina Methylation Arrays | Home Blogs Epigenetic clock, Zymo leading the way Bisulfite Conversion for Illumina Methylation Arrays Troubleshooting & Best Practice GuidelinesMethylation arrays are common platforms for analyzing